The 100k Pathogen Genome Project is lead by the Weimer lab and is a landmark global consortium that seeks to use whole-genome sequencing and metagenomics to examine bacterial diversity as a means to develop methods that are applicable to bacterial identification, functional biomarkers, and source tracking in agriculture, public health, and the environment. The persistence of bacteria in causing disease in humans and animals is of global importance that can be addressed using population genomics concepts based on whole genome sequence and metagenomics
The 100K Pathogen Genome Project is using next-generation and third-generation sequencing approaches to uncover the vast diversity of bacterial genotypes that form the basis of identification and tracking using population genomic approaches that are only recently available to microbiology. Continual bacterial genetic evolution is hindering our ability to consistently detect and mitigate pathogens, which interfere with our preparedness to defend public health. By leveraging genome diversity it enables new diagnostic and public health approaches for the management of diagnostics and phylogeny.
Use of population genetics tools and concepts is a bold and revolutionary approach that is new to microbiology at this scale. Genomics is empowering precise and robust molecular testing that can be used as a forensic tool for many applications across humanity. In spite of extensive efforts to increase regulation and develop early warning diagnostics for improved public health monitoring, definitive interventions remain elusive in large, partly due to the lack of sufficient information about microbial diversity. This effort is aimed at linking diagnostics with functional characteristics encoded in the genome to enable persistence in the environment, mediate zoonotic transmission, and allow host adaptation. The 100K Pathogen Genome Project is focused on tackling these global problems using big data approaches.
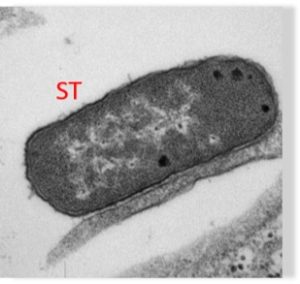
Salmonella infecting a human stem cell (Weimer Lab)
Bart C. Weimer Ph.D., UC Davis School of Veterinary Medicine
1089 Veterinary Medicine Drive, Davis, CA 95616 (530) 752-6426
bcweimerATucdavis.edu, infoAT100kgenomes.org


Заказали тут препарат и остались довольны
венклекста купить